this post was previously published on my old website, there’ll be a few of those older but useful posts that I’ll be migrating over in the next little while…
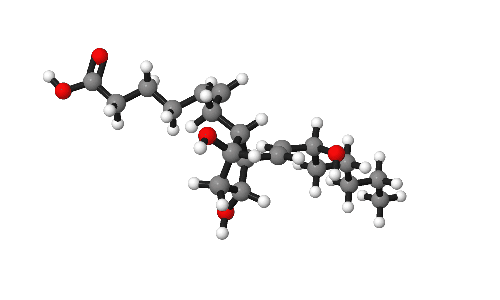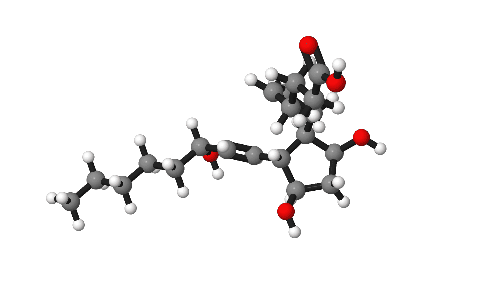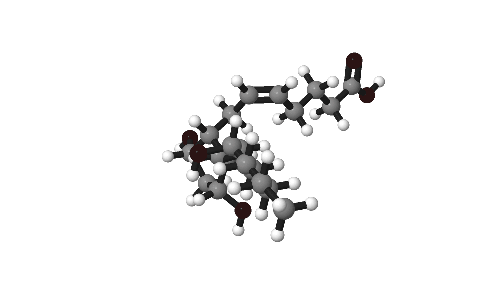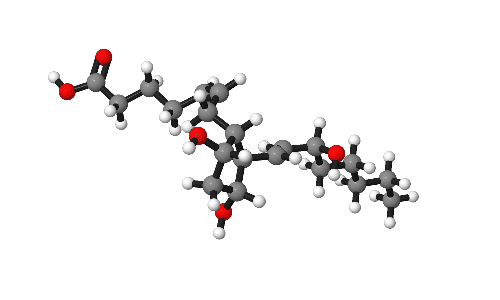
Molecules are beautiful things, intricate and infinitely variable. As part of research publications it can be useful to catch them from their best angles. This short post gives some tips on how to present molecules in publications.
Our model for today is Prostaglandin-F2α